MLatom 3: 10-year anniversary edition is released!
Published Time: 2023-09-22 09:39:06

We are happy to announce that on the occasion of its ten-year anniversary, we have released on September 12 a brand new MLatom 3 with tons of new features including more simulation options, Python API for user-customized workflows, and its integration into XACS cloud computing with the online terminal. You can watch the release broadcast video for an overview of the new release:
Old features stay the same
You can still use MLatom for the same calculations as before – not much changed from outside for the users of the old features, although inside the hoods, we completely rewritten most of MLatom. For example, this input file for geometry optimization works as before:
AIQM1 # or useMLmodel MLmodelIn=CH3Cl.unf if you, e.g., want to use CH3Cl.unf model
xyzfile=init.xyz
optxyz=opt.xyz
geomoptTons of new simulation features
MLatom 3 comes with tons of new useful features such as:
- molecular dynamics
- rovibrational (IR and power) spectra simulations from molecular dynamics
- support of more quantum chemical methods ranging from DFT and ab initio via interfaces to PySCF, Gaussian, and Orca to semi-empirical via interfaces to mndo, Sparrow, and xtb.
- ML-quantum dynamics via an interface to the MLQD package
It means that now you can choose from a huge library of methods and, e.g., use them to run MD via input file as
MD # Molecular dynamics
method=AIQM1 # Use AIQM1
initConditions=user-defined # Use user-defined initial conditions
initXYZ=init.xyz # File with initial geometry
initVXYZ=init.vxyz # File with initial velocities
dt=0.1 # Time step
trun=100000 # Length of trajectory
thermostat=nose-hoover # Use Nose-Hoover thermostat
temperature=300 # Set temperature
trajH5MDout=traj.h5 # Save trajectory in traj.h5
qmprog=mndoPython API
If you feel more comfortable working with Python and Jupyter, you can now use MLatom’s Python API (MLatom PyAPI) to create your own custom workflows by simply importing mlatom and exploiting its flexibility and a wide range of capabilities. For example, if you want to optimize geometry using MLatom PyAPI, your Python script might look like this:
import mlatom as ml # import MLatom module.
ani = ml.models.methods(method='ANI-1ccx') # define an ANI-1cxx model.
init_mol = ml.data.molecule.form_xyz_file('init.xyz') # load the molecular structure.
final_mol = ml.optimize_geometry(model=ani, initial_molecule=init_mol).optimized_molecule # the optimized structure can be obtained by command.
final_mol.xyz_coordinates # we can see the coordinates of the last molecule.
print(final_mol.get_xyz_string()) # print the coordinates in ".xyz" format.
final_mol.write_file_with_xyz_coordnates(filename='final.xyz') # save it in "final.xyz".The documentation for MLatom PyAPI with examples and tutorials will be released soon.
New permissive license
MLatom 3 comes with a new permissive license: MIT (modified to request proper citations in the academic publications).
How to install it?
MLatom 3 can be installed via a variety of way:
pip install mlatom- download the source code and binaries from MLatom website.
- Source code on GitHub:
git clone https://github.com/dralgroup/mlatom.git
More instructions such as installation of dependences will be provided soon in our online manual.
Integration with new XACS cloud computing
MLatom is part of XACS and this allows you to run it in the XACS terminal in your web browser:
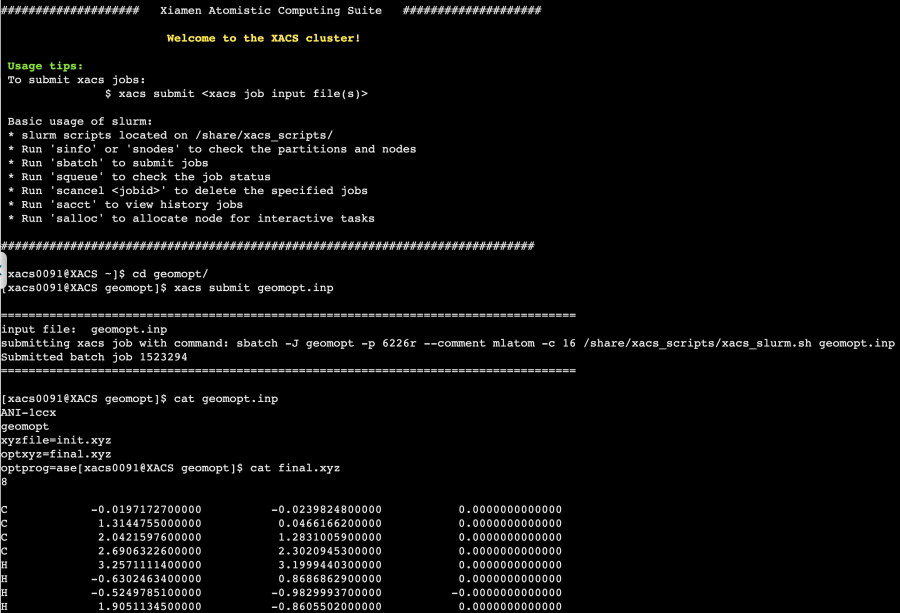
Documentation and support
Please refer to an updated MLatom manual for more information.
While we are continuously improving it with more details and tutorial examples, you might want to join our XACS Support Slack workspace for faster communications and send us your inquiries via email (dral at xmu.edu.cn).
For Chinese users, we also have both the XACS WeChat official channel which you can follow and the user group (send an inquiry to xacs at xmu.edu.cn).